http://www.chemistrymag.org/cji/2003/05a078ne.htm |
Oct. 1, 2003 Vol.5 No.10 P.78 Copyright |
Zeng Xirui, Liu Dongsheng, Peng Shieming#,
Lee Genehsiang#
(Chemistry Department of Jinggangshan Normal College, Ji'an Jiangxi, 343009; # Chemistry Department of National Taiwan
University, Taipei Taiwan)
Keywords solvent effect, complex structure, linear trinuclear copper complex 1. INTRODUCTION
Metal string complexes are mostly used in the fundamental study of metal-metal interactions[1,2] and their potential application as a molecular metal wire. In our past effort, a series of polynuclear metal string complexes have been successfully synthesized and characterized[3-6]. The linear trinuclear copper(II) complex is the simplest metal string that was synthesized early by Brian Hathaway[7] and Gloria J. Pyrka[8], in which the Cu-Cu-Cu bond angle is very close to 180° and the central Cu atom occupies a two-fold special position. Here we report a trinuclear copper string complex that involves solvent molecule in its unit cell, which is apparently different from that of the previous reports due to the effect of the included ether molecule in cell units.
2. EXPERIMENTAL
2.1 Synthesis of the complex
Dpyam (Di-2-pyridylamine, 0.25 g, 1.46 mmol) and CuCl(0.18g, 1.8mmol) were placed in
an Erlemeyer flask, and DMF(25 mL) was added as solvent. The mixture was heated for 10min
at 120-130
2.2 Determination and analysis of the crystal structure
The unit-cell data and intensities were collected on Nonius KappaCCD diffractometer with graphite-monochromated Mo Ka radiation (l = 0.7107 Å). The structures were solved and refined by using the SHELXS-97 and SHELXTL-97 package, respectively. An empirical absorption correction based on three azimuthal scans was also applied. The structures were solved using direct methods and difference Fourier techniques and refined by least-squares analysis. The non-hydrogen atoms were refined anisotropically, and the hydrogen atoms were included in an idealized geometry but not refined. The crystal data are summarized in Table 1. In which Hathaway's data about compound [Cu3(m3-dpa)4Cl2·2H2O] have been cited[7] and compared with our experimental data about compound [Cu3(m3-dpa)4Cl2·ether].
Table 1 Crystal data and structure refinement for two complexes
Empirical formula |
[Cu3( m3-dpa)4Cl2·ether] |
[Cu3( m3-dpa)4Cl2·2H2O] |
Formula weight |
1016.42 |
960.31 |
Crystal system |
Monoclinic |
orthorhombic |
Space group |
P2(1)/c |
Pnn2 |
Unit cell dimensions |
a = 16.0093(1) Å a=90°b = 15.7496(1) Å b=99.22 (4)° c = 17.0157(1) Å g=90° |
a = 14.092(3) Åb = 12.895(3) Å c = 11.190(2) Å |
Volume, Z |
4234.90(5) Å3 , 4 |
2031.0 Å3 , 2 |
Density (calculated) |
1.594 Mg/m3 |
1.57 Mg/m3 |
Absorption coefficient |
1.671 mm-1 |
1.678 mm-1 |
F(000) |
2076 |
947 |
Independent reflections |
9734 [R(int) = 0.0660] |
|
Reflections collected |
33461 |
1873 |
Final R indices[I>2sigma(I)] |
R1 = 0.0395, wR2 = 0.0892 |
R1 = 0.028, wR2 = 0.041 |
Largest diff. peak and hole |
0.784 and -0.903 e. Å-3 |
0.58 and -0.37e. Å-3 |
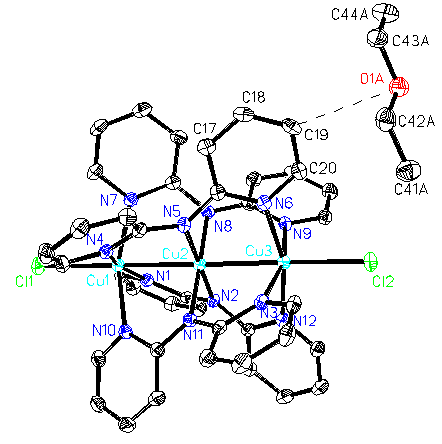 |
Fig. 1 ORTEP plot with 50% probability ellipsoids of [Cu3(dpa)4Cl2]·ether, perpendicular to the Cl-Cu-Cu-Cu-Cl axis. |
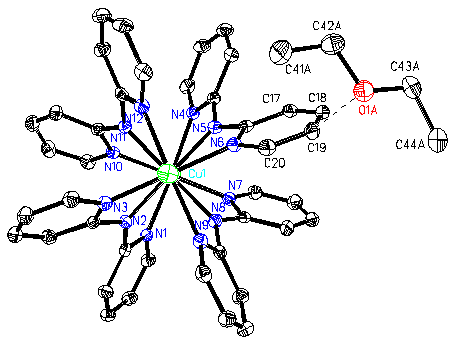 |
Fig. 2 ORTEP plot with 50% probability ellipsoids of [Cu3(dpa)4Cl2]·ether, along the Cl-Cu-Cu-Cu-Cl axis. |
3. RESULTS AND DISCUSSION
The structure of the complex [Cu3(m3-dpa)4Cl2 {dpa=di-2-pyridylamino,
[(C5H4N)2N]-}] is shown in Fig. 1
(perpendicular to the Cl-Cu-Cu-Cu-Cl axis) and Fig. 2 (along the copper-copper axis), both
included solvent ether molecule in unit cell. As other metal string complexes consisted of
polypyridylamido ligand, the three CuII ions and the two chloride ions are
collinear with all the angles of M-M-M being ~180° and the linear metal chain is
wrapped by four all-syn of deprotonated oligo-a-pyridylamido ligands. The mean central Cu-N distances are below
2.0 Å, while the mean terminal Cu-N
distances are greater than 2.0 Å(see Table 2).
These data are similar to the previous descriptions.
By scrutinizing, we find that the Cu(1)-Cu(2) and Cu(2)-Cu(3)
distances, with 2.4691(5) and 2.4774(5) Å
respectively, are not completely equal, Dd=0.0083 Å, this value has gone beyond normal error range, furthermore the
Cu-Cl lengths in both ends of metal string, with 2.4404(8) and 2.4740(8) Å respectively,
have more evident differences. Selected geometric parameters have been listed in Table 2.
Table 2 Selected bond distances (
Å) and angles (°)Cu(1)-Cu(2) |
2.4691(5) |
Cu(2)-Cu(3) |
2.4774(5) |
Cu(1)-Cl(1) |
2.4404(8) |
Cu(3)-Cl(2) |
2.4740(8) |
Cu(1)-N(1) |
2.065(3) |
Cu(2)-N(2) |
1.962(2) |
Cu(3)-N(3) |
2.076(3) |
O(1)-Cu(3) |
6.594 |
Cu(1)-Cu(2)-Cu(3) |
178.94(2) |
N(1)-Cu(1)-Cu(2) |
81.99(7) |
N(2)-Cu(2)-Cu(1) |
90.29(7) |
N(3)-Cu(3)-Cu(2) |
81.49(7) |
Cl(1)-Cu(1)-Cu(2) |
177.56(3) |
Cl(2)-Cu(3)-Cu(2) |
178.95(3) |
It is clear that the complex molecule structure has deviated from crystallographic twofold symmetry described in Hathaway and Pyrka's paper. We deduce that the solvent molecule involved in the unit cell resulted in the change of the complex molecular geometry. As shown in Fig.1, solvent ether molecule is lying on Cu(3) side of the Cl(1)-Cu(1)-Cu(2)-Cu(3)-Cl(2) chain and Cu(3)-O(1A) distance is 6.58
Å. It is noteworthy that intermolecular hydrogen bonds would impact on the geometry of the complex. First, the second ether molecule would elongate bond length of Cu(2)-Cu(3) lying on this side through hydrogen bond C(19)-H(19A)-O(1A) with distance D-A 3.326(4) Å [here, the O(1A) is an atom from adjacent symmetry-related unit]. Second, two terminal Cl atoms are taken as acceptor and form intermolecular hydrogen bonds C(2)-H(2)-Cl(1) and C(39)-H(39A)-Cl(2), respectively [also, the C(2), H(2), C(39) and H(39A) are atoms from adjacent symmetry-related unit], the unequals on hydrogen bonds action would result in the difference of bond lengths Cu(1)-Cl(1) and Cu(3)-Cl(2) on two ends, the multi-hydrogen bonds would consequently affect the structure symmetry of the complex Cu3(m3-dpa)4Cl2. Related data are listed in Table 3. Similar cases can be found in compounds [Co3(m3-dpa)4Cl2·CH2Cl2], [Co3(m3-dpa)4Cl2·2CH2Cl2] and [Co3(m3-dpa)4Cl2·2CH2Cl2·H2O] [9,10], the first one is symmetric geometry and the latter two are unsymmetric structures.The difference of involved solvent molecule in unit cell also resulted in the difference of the crystal cell parameters. Related data can be examined in Table 1, in which, the data of complex [Cu3(m3-dpa)4Cl2·2H2O] are Hathaway's results7. It is obvious that many parameters such as crystal system, space group, unit cell dimensions and so on are different, by this token, it is not suitable to distinguish crystal structure just by cell parameters.
Table 3. Related hydrogen-bonding geometry(
Å, °)D- H··· A |
D- H |
H ··· A |
D---A |
D- H ··· A |
C(2)- H(2) ··· Cl(1) |
0.95 |
2.80 |
3.681(3) |
154 |
C(39)- H(39A) ··· Cl(2) |
0.95 |
2.76 |
3.676(3) |
162 |
C(19)- H(19) ··· O(1) |
0.95 |
2.52 |
3.326(4) |
143 |
4. CONCLUSION
The solvent species and the amount involved in crystal unit cell would result in the
difference of the crystal cell parameters. Also, the complex structure and geometry maybe
changed in varying degree because of the interaction between complex and solvent molecules
through the hydrogen bonds and Van der Waals forces.
REFERENCES
[1] Yang M H, Lin T W, Peng S M et al. Chem. Commun., 1997, 2279.
[2] Cotton F A, Daniels L M, Murillo C A et al. Chem. Commun., 1998, 39.
[3] Sheu J T, Liu C C, Peng S M et al. Chem. Commun., 1996, 315.
[4] Shieh S J, Chou C C, Peng S M et al. Angew. Chem.Int. Ed. Engl., 1997, 36, 56.
[5] Chang H C, Lin T W, Peng S M. European Journal of Inorganic Chemistry, 1999, 1243.
[6] Peng S M, Wang C C, Jang Y L et al. J. Magnet. & Magnet. Mater., 2000, 209, 80.
[7] Wu L P, Field P, Hathaway B. J. Chem. Soc. Dalton Trans., 1990, 3835.
[8] Pyrka G J, EI-Mekki M, Pinkerton A A. J. Chem. Soc., Chem. Commun., 1991, 84.
[9] Yang E C, Cheng M C, Peng S M et al. J. Chem. Soc., Chem. Commun., 1994, 2377.
[10] Cotton F A, Daniel G T et al. Chem. Commun. 1997, 421.