http://www.chemistrymag.org/cji/2005/072017ne.htm |
Feb.1, 2005 Vol.7 No.2 P.17 Copyright |
Synthesis and structure of the new complex [Cu(ipn)(phen)Cl]Cl·(H2O)2
Du Guixiang, Li Jun, Yang Lining, Wang Wei,
Li Hengxin, Zhang Fengxing
(Department of Chemistry, Northwest University, Shaanxi Key Laboratory of
Physico-Inorganic Chemistry, Xi'an 710069, China)
Supported by The Natural Science Foundation of Shaanxi province (No.HF0125) and the Education Commission Foundation of Shaanxi province (No.01JK078)
Abstract The complex [Cu(ipn)(phen)Cl](Cl)·(H2O)2 (ipn=1,2-diaminopropane, phen=1,10-phenanthroline) has been synthesized and structurally characterized. It crystallizes in the triclinic system, space group P2/c with a=0.7568(7) nm, b=1.541(15) nm, c=1.652(16) nm, a=97.247o, b=91.502o, g=103.933o, V=1.853(3) nm3, wR2=0.1661 and Z=2. The diffraction data (I>2¦Ò (I)) is 4644. The copper ion has a slightly distorted square pyramidal configuration with two nitrogen atoms from phen, two nitrogen atoms from ipn, and one chloride anion.Keywords Copper complex, crystal structure, synthesis. The penta-coordinated copper (II) complexes comprising of both molecules of bidentate diimine, 2,2'-bipyridine (bipy) and 1,10-phenanthroline (phen), and one monodentate co-ligand have been attracting great interests for their diverse stereo and physicochemical properties[1-3]. The molecular structures of five-coordinate copper (II) complexes show an extensive variability, ranging from regular trigonal bipyramidal to regular square based pyramidal with most complexes displaying a structure that is intermediate between the two extreme stereochemistries[4-5]. Although many five-coordinated copper (II) complexes are well documented, to the best of our knowledge, the presence of a mixed ligand compound has been rarely observed[3] and the examples of CuIIN4X (X=Cl-, Br-, I-) complexes generated from mixed ligands are scant[6]. In this paper, we describe the synthesis and crystal structure of a new complex contaning CuIIN4Cl chromophores, [Cu(ipn)(phen)Cl]Cl·(H2O)2.
1. EXPERIMENTAL
1.1 Materials and physical measurements
All reagents were of analytical grade from commercial sources and were used without any further purification. The C, H and N elemental analyses were performed using a Vario EL-III instrument. Infrared spectrum of the title complex was recorded with Equinox 55 spectrophotometer in the 4000--400 cm-1 region using a powdered sample on a KBr plate. The diffraction data were collected on a Brucker SMART APEX II CCD diffractometer using graphite monochromated MoKa radiation (l=0.071073 nm).
1.2 Preparation of [Cu(ipn)(phen)Cl](Cl)(H2O)
Copper dichloride dihydrate (0.170g, 1.0mmol) dissolved in 20mL of water was allowed to react with 20mL of methanol solution of phenanthroline (0.198g, 1.0mmol). Into the resulting solution was poured slowly 10 mL of methanol solution of 1,2-diaminopropane (0.074g, 1.0mmol), the entire mixture was stirred for 1h and then allowed to evaporate slowly in an open atmosphere. After a few weeks, blue blocks of the title complex suitable for X-ray structure study were obtained. The crystal was separated, washed with ethanol, and dried in the air. Yield: 0.316g (74.38%). Anal.calcd.(found) for C15H22Cl2N4O2Cu: C, 42.41 (42.28); H, 5.22 (5.15); N, 13.19 (13.05)%. IR (KBr, cm-1): 3377[n (NH2)], 3122[n(H2O)], 1588,1517[n (phen)].
1.3 X-ray crystallography
Diffraction data were collected by w-2q scan technique. The total reflections of 6419 were collected, of which independent reflections of 4644 could be observed and used in the structure refinement. The structure was solved by direct methods. The positions of all hydrogen atoms were obtained from successive Fourier syntheses. The positions of all non-hydrogen atoms were refined anisotropically with full-matrix least-squares on F2. In the final difference map, the residuals are 1363 and -497 e/nm3, respectively.
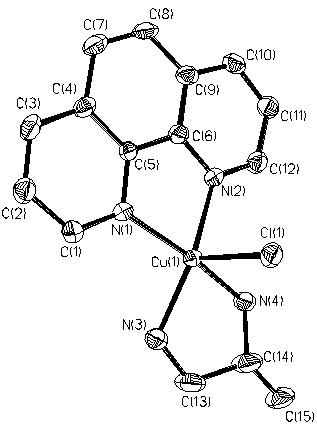
Fig.1 ORTEP projection of [Cu (ipn)(phen)Cl]+ 2. RESULTS AND DISCUSSION
Figure 1 shows the ORTEP projection of the cation [Cu(ipn)(phen)Cl]+. Figure 2 shows one-dimension chain structure of the title complex. The crystallographic data and analysis parameters are showed in Table 1. Selected bond lengths and bond angles are listed in Table 2.
Table 1 Crystal data and structure analysis parameters
complex |
[Cu(ipn)(phen)Cl](Cl)·(H2O)2 |
V (nm3) |
1.853(3) |
Crystal size (mm3) |
0.40¡Á0.35¡Á0.28 |
Z |
2 |
Empirical formula |
C15H22Cl2N4O2Cu |
Dc (g/cm3) |
1.523 |
Temperature (K) |
298(2) |
F (000) |
876 |
Wavelength (nm) |
0.071073 |
Scan mode |
¦Ø-2¦È |
Formula weight |
424.80 |
q range for data collection (o) |
1.72-25.00 |
Crystal system |
triclinic |
Absorption coefficient (mm-1) |
1.482 |
Space group |
P2/c |
Reflections collected |
6419 |
a (nm) |
0.7568(7) |
Independent reflections |
4644 |
b (nm) |
1.541(15) |
Goodness-of-fit |
1.055 |
c (nm) |
1.652(16) |
R [I>2s (I)] |
0.0517 |
a(o) |
97.247 |
wR2 |
0.1661 |
b(o) |
91.502 |
Maximum diff. peak (e/nm3) |
1363 |
| g(o) | 103.933 |
Minimum diff. peak (e/nm3) |
-497 |
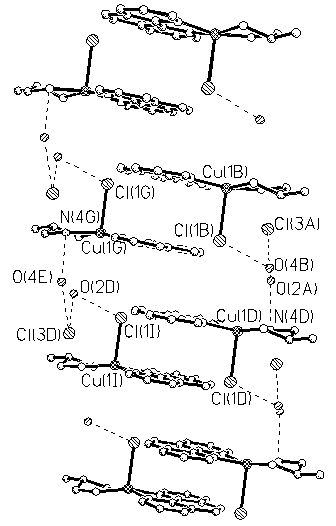
Fig.2 One-dimension chain framework of the title complex
Table 2 Selected bond lengths (nm) and bond angles (o)
Bond lengths |
|||
Cu(1)-N(1) |
0.2024(3) |
Cu(1)-N(4) |
0.2009(3) |
Cu(1)-N(2) |
0.2042(3) |
Cu(1)-Cl(1) |
0.2556(11) |
Cu(1)-N(3) |
0.2001(3) |
||
Bond angles |
|||
N(3)-Cu(1)-N(4) |
83.12(13) |
N(1)-Cu(1)-N(2) |
81.35(12) |
N(3)-Cu(1)-N(1) |
95.05(13) |
N(3)-Cu(1)-Cl(1) |
99.26(11) |
N(4)-Cu(1)-N(1) |
166.29(14) |
N(4)-Cu(1)-Cl(1) |
95.38(11) |
N(3)-Cu(1)-N(2) |
167.35(14) |
N(1)-Cu(1)-Cl(1) |
98.32(9) |
N(4)-Cu(1)-N(2) |
97.48(13) |
N(2)-Cu(1)-Cl(1) |
93.27(9) |
¡¡
The cation [Cu(ipn)(phen)Cl]+ of the title complex possesses five-coordinated environment with CuIIN4Cl chromophores, namely, the Cu atom is coordinated by four nitrogen atoms from one bidentate diimine (phen) and one bidentate diamine (ipn) ligand with one chloride anion forming weak Cu-Cl apex bond. The Cu-N bond lengths and the N-Cu-N bond angles (see Table 2) are similar to the previous complex [Cu(en)(bipy)Cl][OH]
·(H2O)3[6]. The Cu-Cl distance (0.2556(11) nm) is similar to that in the common CuIIN4Cl complexes[7-11], and shorter than the complex [Cu(phen)2Cl][OH](CH3OH)2·(H2O)6[6]. Furthermore, the largest bond angles around the Cu(II) center (b: N(3)-Cu(1)-N(2)=167.35(14)¡ã) is slightly larger than the second-largest one (¦Á: N (4)-Cu(1)-N(1)=166.29¡ã). The value of the differences between the longest and shortest Cu-N bond distances DCu-N is 0.0041nm. These imply that the geometry around the copper (II) center in the title complex is of distorted square-pyramidal one. Since the angular structural index parameter, t=(b-a)/60, is evaluated by the two largest angles (a<b) in the five-coordinate geometry[12], the value of t=0.017 indicates a regular square based pyramidal (t value of (b-a)/60 is defined, where t=1.0 for a regular trigonal bipyramidal stereochemistry and t=0.0 for a regular square based pyramidal stereochemistry). Hydrogen bonds play an important role in the crystal packing and the stability of the inclusion complex (Fig.2.). Adjacent units are cross-linked by three types of intermolecular hydrogen bonds, which stabilize the conformation with the distances of 0.232nm for (H2O) O¡Cl, 0.221nm for (H2O) O¡H-N(ipn), 0.240nm for Cl¡H-N(ipn). 1,10-phenanthroline rings in the [Cu (ipn)(phen)Cl]+ cation are parallel to each other, which produce p-p stacking interactions as shown in Fig.2, and the interplanar distance is 0.3143nm. A novel one-dimension chain was formed through hydrogen bonding and p-p stacking interactions.REFERENCES
[1] Tran D, Skelton B W, White A H et al. Ford P.C. Inorg. Chem. 1998, 37. 2505-2511.
[2] Nagle P, Sullivan E O', Hathaway B J et al. J. Chem. Soc. Dalton Trans. 1990,
3399-3406.
[3] Su C C, Lin Y L, J S et al. Polyhedron. 1993, 12. 2687-2696.
[4] Hathaway B J, Billing D E. Coord Chem Rew. 1970, 5. 143-167.
[5] Hathaway B J, Struct Bonding (Berlin) 1984, 57. 55.
[6] Mao H Y, Shen X Q, Li G et. al., Polyhedron. 2004, 23. 1961-1967.
[7] Sullivan C O¡¯, Murphy G, Murphy B et al. Chem.
Soc. Dalton Trans. 1999, 1835-1844.
[8] Brophy M, Murphy G, Sullivan C O et al. Polyhedron. 1999, 18. 611-615.
[9] Murphy M, Nagle P, Murphy B et al. Chem.Soc. Dalton Trans. 1997, 2645-2652.
[10] O¡¯Brien E C, Farkas E, Gil M J et al. J. Inorg.
Biochem. 2000, 79. 47-51.
[11] Ma B.Q, Gao S, Yi T, et al. Inorg. Chem. Commun. 2000, 3. 93-95.
[12] Addison A W, Ral T N, Reedijk J et al. J. Chem.Soc. Dalton Trans. 1984, 1349-1356.
ÅäºÏÎï[Cu(ipn)(phen)Cl]Cl·(H2O)2µÄºÏ³ÉºÍ¾§Ìå½á¹¹
¶Å¹ðÏã Àî¬B ÑîÀòÄþ Íõΰ ÀîºãÐÀ
ÕÅ·êÐÇ
£¨ÉÂÎ÷Ê¡ÎïÀíÎÞ»ú»¯Ñ§ÖصãʵÑéÊÒ£¬Î÷±±´óѧ»¯Ñ§Ïµ£¬Î÷°² 710069£¬Öйú£©
ÕªÒª ±¾ÎÄÒÔÁڷȆªßø(phen)ºÍ1,2-±û¶þ°·(ipn)ΪÅäÌåÖƵÃÁËÐÂÅäºÏÎï[Cu(ipn)(phen)Cl]Cl(H2O)2,²â¶¨ÁËÆ侧Ìå½á¹¹¡£¸Ãµ¥¾§ÊôÈýб¾§Ïµ£¬¿Õ¼äȺΪP2/c£¬¾§°û²ÎÊýa=0.7568(7)nm£¬b=1.541(15)
nm,c=1.652(16)nm,a=97.247o,b=91.502o,g=103.933o,V=1.853(3) nm3,wR2=0.1661£¬Z=2¡£ÑÜÉäÊý¾Ý(I>2s (I))Ϊ4644¡£¾§Ìå½á¹¹±íÃ÷£¬ÅäºÏÎïÖÐCu2£«´¦ÓÚÒ»¸öÉÔ΢ŤÇúµÄËĽÇ׶Åäλ»·¾³£¬ÅäλÔ×Ó·Ö±ðÀ´×ÔÁڷȆªßøµÄÁ½¸öµªÔ×Ó£¬1,2-±û¶þ°·µÄÁ½¸öµªÔ×ÓºÍÒ»¸öÂÈÀë×Ó¡£
¹Ø¼ü´Ê ÍÅäºÏÎ¾§Ìå½á¹¹£¬ºÏ³É